A demo of K-Means clustering on the handwritten digits data¶
In this example with compare the various initialization strategies for K-means in terms of runtime and quality of the results.
As the ground truth is known here, we also apply different cluster quality metrics to judge the goodness of fit of the cluster labels to the ground truth.
Cluster quality metrics evaluated (see Clustering performance evaluation for definitions and discussions of the metrics):
| Shorthand | full name |
|---|---|
| homo | homogeneity score |
| compl | completeness score |
| v-meas | V measure |
| ARI | adjusted Rand index |
| AMI | adjusted mutual information |
| silhouette | silhouette coefficient |
Script output:
n_digits: 10, n_samples 1797, n_features 64
_______________________________________________________________________________
init time inertia homo compl v-meas ARI AMI silhouette
k-means++ 1.47s 69432 0.602 0.650 0.625 0.465 0.598 0.146
random 1.40s 69694 0.669 0.710 0.689 0.553 0.666 0.147
PCA-based 0.11s 71820 0.673 0.715 0.693 0.567 0.670 0.150
_______________________________________________________________________________
Python source code: plot_kmeans_digits.py
print __doc__
from time import time
import numpy as np
import pylab as pl
from sklearn import metrics
from sklearn.cluster import KMeans
from sklearn.datasets import load_digits
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
np.random.seed(42)
digits = load_digits()
data = scale(digits.data)
n_samples, n_features = data.shape
n_digits = len(np.unique(digits.target))
labels = digits.target
sample_size = 300
print "n_digits: %d, \t n_samples %d, \t n_features %d" % (n_digits,
n_samples, n_features)
print 79 * '_'
print ('% 9s' % 'init'
' time inertia homo compl v-meas ARI AMI silhouette')
def bench_k_means(estimator, name, data):
t0 = time()
estimator.fit(data)
print '% 9s %.2fs %i %.3f %.3f %.3f %.3f %.3f %.3f' % (
name, (time() - t0), estimator.inertia_,
metrics.homogeneity_score(labels, estimator.labels_),
metrics.completeness_score(labels, estimator.labels_),
metrics.v_measure_score(labels, estimator.labels_),
metrics.adjusted_rand_score(labels, estimator.labels_),
metrics.adjusted_mutual_info_score(labels, estimator.labels_),
metrics.silhouette_score(data, estimator.labels_,
metric='euclidean',
sample_size=sample_size),
)
bench_k_means(KMeans(init='k-means++', k=n_digits, n_init=10),
name="k-means++", data=data)
bench_k_means(KMeans(init='random', k=n_digits, n_init=10),
name="random", data=data)
# in this case the seeding of the centers is deterministic, hence we run the
# kmeans algorithm only once with n_init=1
pca = PCA(n_components=n_digits).fit(data)
bench_k_means(KMeans(init=pca.components_, k=n_digits, n_init=1),
name="PCA-based",
data=data)
print 79 * '_'
###############################################################################
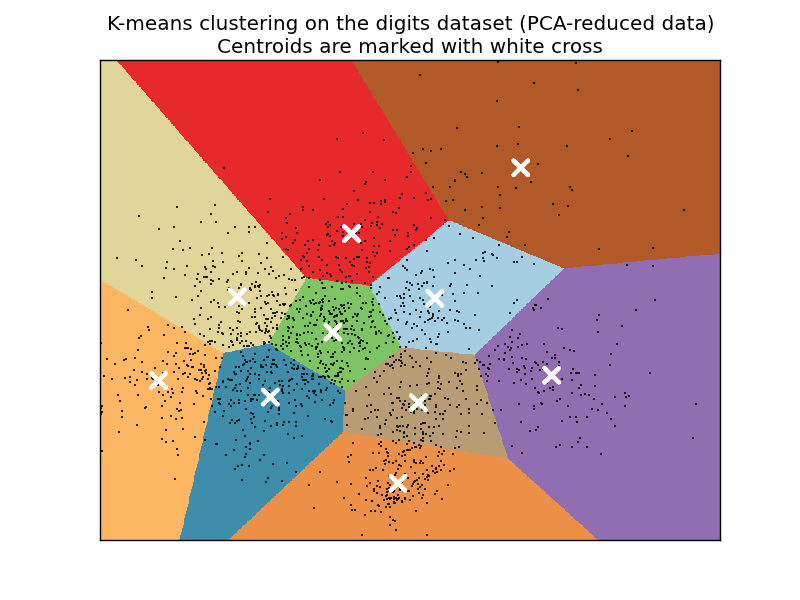
# Visualize the results on PCA-reduced data
reduced_data = PCA(n_components=2).fit_transform(data)
kmeans = KMeans(init='k-means++', k=n_digits, n_init=10).fit(reduced_data)
# Step size of the mesh. Decrease to increase the quality of the VQ.
h = .02 # point in the mesh [x_min, m_max]x[y_min, y_max].
# Plot the decision boundary. For that, we will asign a color to each
x_min, x_max = reduced_data[:, 0].min() + 1, reduced_data[:, 0].max() - 1
y_min, y_max = reduced_data[:, 1].min() + 1, reduced_data[:, 1].max() - 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# Obtain labels for each point in mesh. Use last trained model.
Z = kmeans.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
pl.figure(1)
pl.clf()
pl.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()),
cmap=pl.cm.Paired,
aspect='auto', origin='lower')
pl.plot(reduced_data[:, 0], reduced_data[:, 1], 'k.', markersize=2)
# Plot the centroids as a white X
centroids = kmeans.cluster_centers_
pl.scatter(centroids[:, 0], centroids[:, 1],
marker='x', s=169, linewidths=3,
color='w', zorder=10)
pl.title('K-means clustering on the digits dataset (PCA-reduced data)\n'
'Centroids are marked with white cross')
pl.xlim(x_min, x_max)
pl.ylim(y_min, y_max)
pl.xticks(())
pl.yticks(())
pl.show()
